The eigenvalues or singular values provide information about the amount of variance captured by each principal component. Step 6: Determining the Captured Variance The loadings, represented by the eigenvectors, indicate how much each experiment contributes to each principal component. These principal components represent the directions of maximal variance in the data. The principal components are obtained by multiplying the mean-subtracted data matrix by the eigenvectors. Step 5: Finding the Principal Components and Loadings Eigenvalues are also computed, representing the amount of variance captured by each eigenvector. These eigenvectors represent the directions in which the data exhibits the maximum variance. 
The leading eigenvectors of the covariance matrix are computed. Step 4: Computing the Eigenvectors and Eigenvalues It is computed as the transpose of the mean-centered data matrix multiplied by the mean-centered data matrix. The covariance matrix represents the correlations between the rows of the data matrix. The covariance matrix is computed using the mean-centered data matrix. To bring the center of the data distribution to the origin, the mean vector is subtracted from the data matrix. This mean vector represents the average row values and is denoted as X bar. The mean of the data matrix is computed by averaging the rows. To compute PCA using SVD, the following steps are involved: Step 1: Computing the Mean 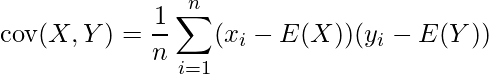
In order to model this data, the mean is computed and subtracted from the data matrix, resulting in zero-mean data. The data is assumed to have a statistical distribution with some variability. In PCA, the data matrix is represented as a transpose, with each row vector representing measurements from a single experiment. There are some differences in notation and conventions between the PCA and SVD literature. 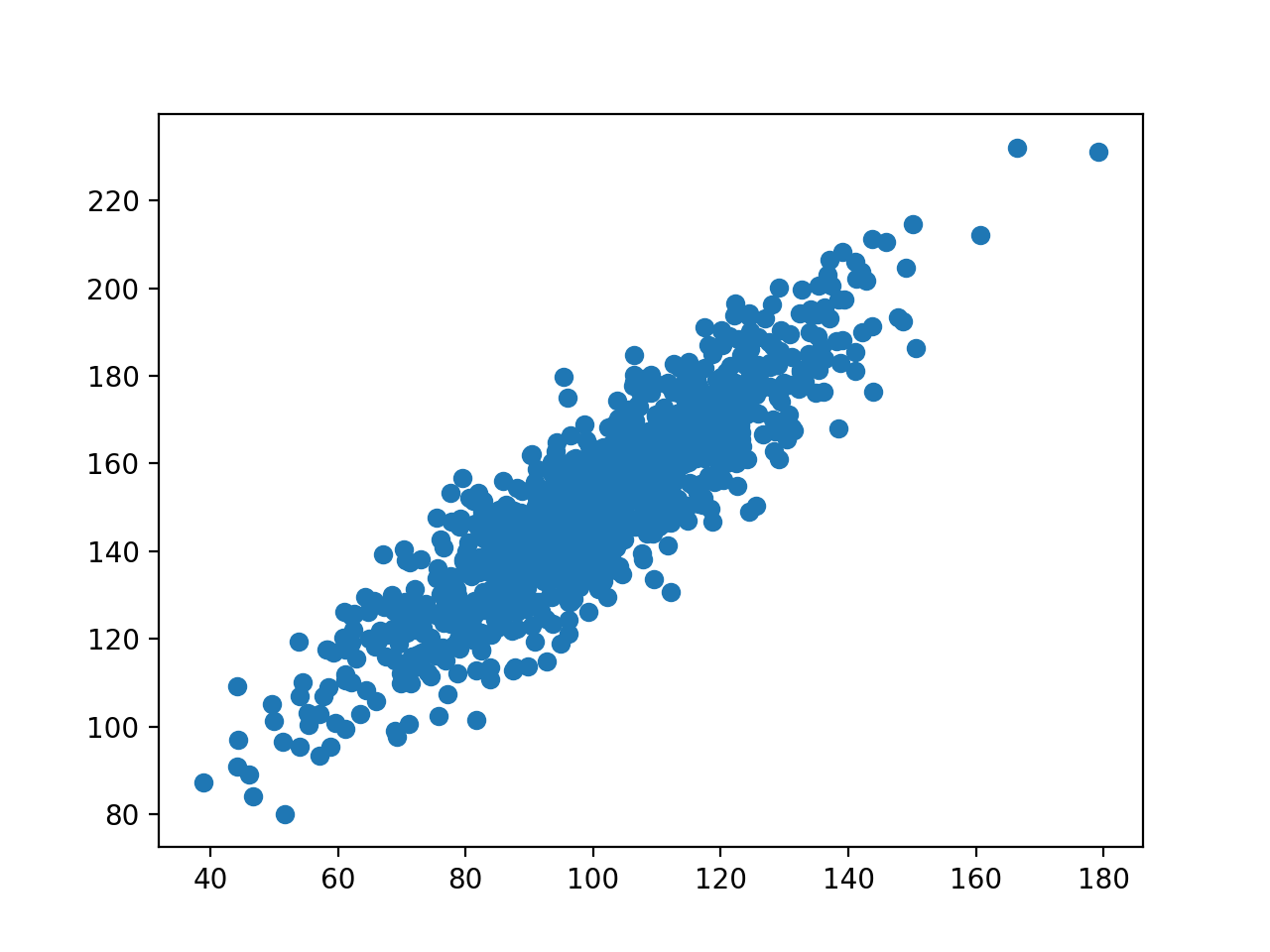
The statistical interpretation of SVD forms the basis for PCA. The Statistical Interpretation of Singular Value Decomposition (SVD) By doing so, it provides a hierarchical coordinate system based on the directions in the data that capture the statistical variations. PCA aims to uncover the dominant combinations of features that describe the maximum amount of variance in a data set. It is a well-established method with a strong theoretical foundation. PCA has a long history and has been widely used since its introduction in 1901 by Pearson. Background of Principal Component Analysis (PCA) Additionally, we will examine the concept of captured variance and the visualization of PCA results. We will discuss the steps involved in PCA computation, implementation in MATLAB, and provide examples using real data sets. In this article, we will explore how to compute PCA using Singular Value Decomposition (SVD), focusing on the statistical interpretation of the SVD. It is widely applied in data science and machine learning applications to uncover low-dimensional Patterns and build models. Principal Component Analysis (PCA) is a fundamental technique used in probability and statistics to reduce the dimensionality of data. Example 2: Genetic Markers for Ovarian Cancer.Step 6: Determining the Captured Variance.Step 5: Finding the Principal Components and Loadings.Step 4: Computing the Eigenvectors and Eigenvalues.Step 3: Computing the Covariance Matrix.The Statistical Interpretation of Singular Value Decomposition (SVD).Background of Principal Component Analysis (PCA).Wolfram Language & System Documentation Center.Discover Low-Dimensional Patterns with Principal Component Analysis "Covariance." Wolfram Language & System Documentation Center. Wolfram Research (2007), Covariance, Wolfram Language function, (updated 2023). Cite this as: Wolfram Research (2007), Covariance, Wolfram Language function, (updated 2023). 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
